In this section, we discuss hypothesis tests for two population standard deviations. In other words, we discuss methods of inference for the standard deviations of one variable from two different populations. These methods are based on the $F$-distribution, named in honour of Sir Ronald Aylmer Fisher.
The $F$-distribution is a right-skewed probability density distribution with two shape parameters, $v_{1}$ and $v_{2}$, called the degrees of freedom for the numerator ($v_{1}$) and the degrees of freedom for the denominator ($v_{2}$).
$$ df = (v_{1}, v_{2})$$As for any other density curve, the area under the curve of the $ F$ distribution corresponds to probabilities. The area under the curve and, thus, the probability for any given interval and given $df$ is computed with software. Alternatively, one may look them up in a table. In those tables, generally, the degrees of freedom for the numerator ($v_{1}$) are displayed along the top, whereas the degrees of freedom for the denominator ($v_{2}$) is displayed in the outside column on the left.
In order to perform a hypothesis test for two population standard deviations, the value corresponding to a specified area under a $F$-curve is calculated.
Given $\alpha$, where $\alpha$ corresponds to a probability between 0 and 1, $F_{\alpha}$ denotes the value having an area $\alpha$ to its right under a $ F$ curve.
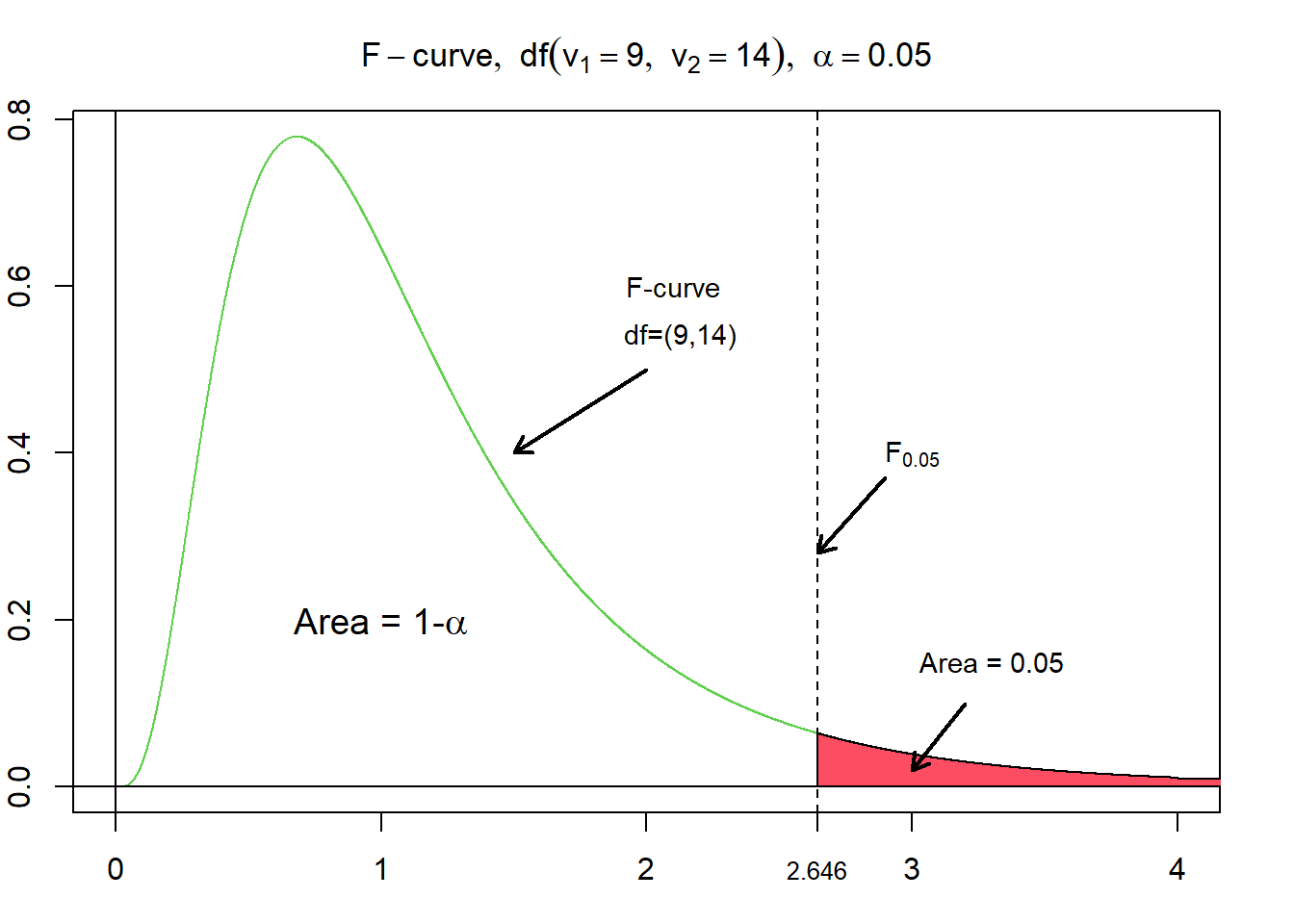
The figure above illustrates the probability density function of a F distribution with the degrees of freedom of 9 and 14. Additionally, the rejection and non-rejection areas regarding a right-tailed hypothesis test are added to a significance level of 95 %, respectively, with an alpha level of 5%. The corresponding, critical value, of $F_{0.05}$ for $df = (9,14)$ evaluates to $\approx 2.6458$.
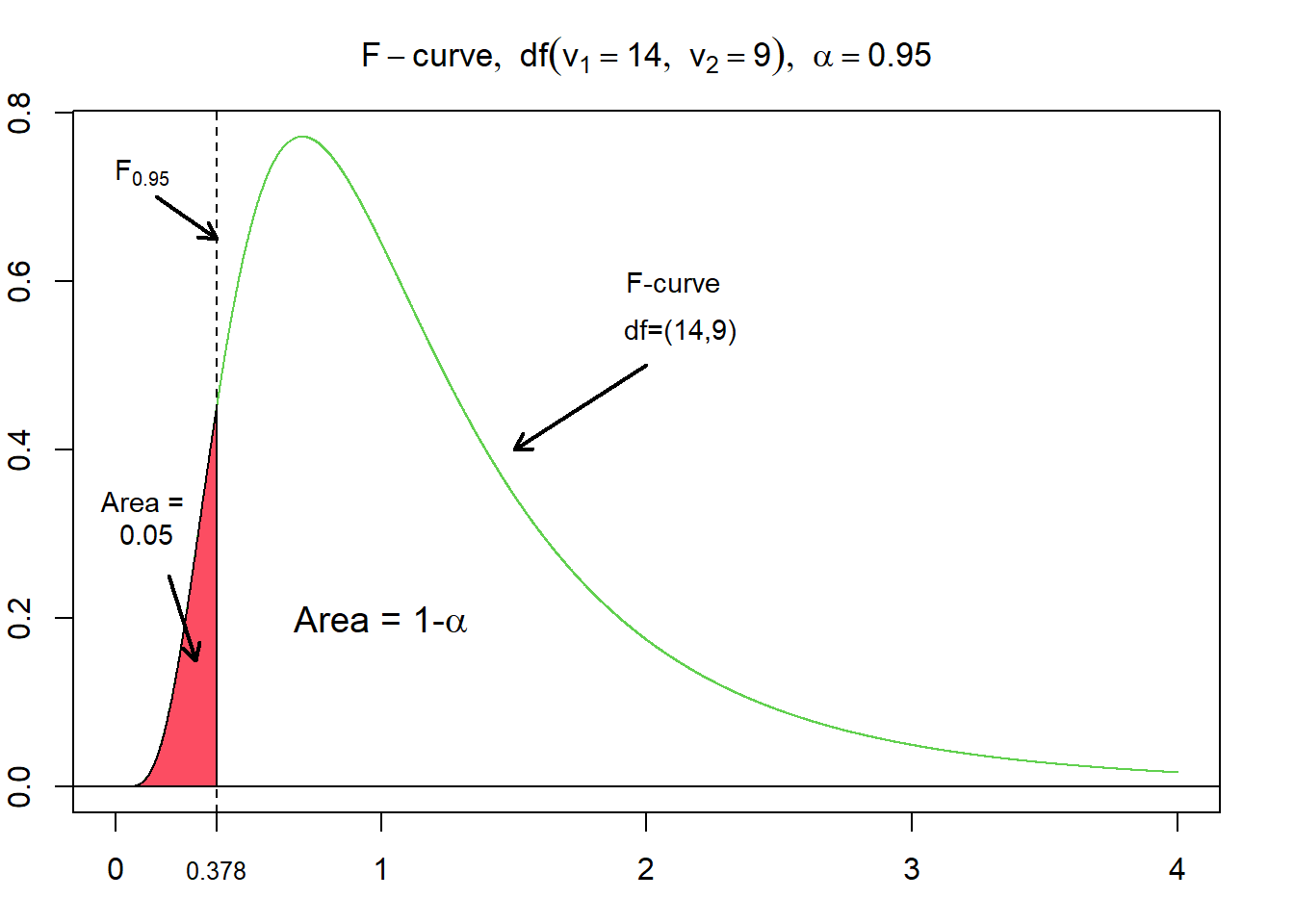
One interesting property of $F$-curves is the reciprocal property. This means, that for a $F$-curve with $df = (v_{1}, v_{2})$, the $F$-value having the area $\alpha$ to its left equals the reciprocal of the $F$-value having the area $\alpha$ to its right for an $F$-curve with $df = (v_{2}, v_{1})$ (Weiss, 2010). Adopted to the example from above, where $F_{0.05}$ for $df=(9,14)$ evaluates to $\approx 2.6458$, this means that $F_{0.95}$ for $df=(14,9)$ evaluates to $\frac {1} {2.6458} = 0.378$.
Interval Estimation of $\sigma_{1} - \sigma_{2}$¶
The $100(1 - \alpha)\ \%$ confidence interval for $\sigma$ is
$$\frac {1} {\sqrt {F_{\alpha /2} } } \times \frac {s_{1}} {s_{2}} \le \sigma \le \frac {1} {\sqrt {F_{1 - \alpha /2} } } \times \frac {s_{1}} {s_{2}} \text{,} $$where $s_1$ and $s_2$ are the sample standard deviations.
